Roles of H3K27me3 Demethylase JMJD3 in Inflammation and Cancers
Xia Chen1, Xue Xiao2, Fei Guo3*
1Department of Gynecology and Obstetrics, The First Affiliated Hospital of Nanchang University, Nanchang, China
2Anesthesiology of the Second Clinical Medical College, Nanchang University
3Burn Center, The First Affiliated Hospital of Nanchang University, Nanchang, China
Abstract
Histone demethylation is an important part of epigenetic modifications, involving in multiple physiological and pathophysiological processes such as proliferation, differentiation, senescence, apoptosis, reprogramming and so on. JmjC domain-containing protein D3 (JMJD3, also called KDM6B) specifically demethylates lysine 27 on histone H3 (H3K27me3), a repressive epigenetic mark, therefore modulating the expression of target genes. JMJD3 can be strongly and quickly induced by various inflammatory stimuli and cellular stresses, and can enhance pro-inflammatory reactions as well as anti-inflammatory reactions by targeting diverse transcription factors in gene promoters and bodies. Additionally, JMJD3 has a dual effect on many types of cancers through binding to promoters of oncogenes or suppressor genes. As is known to us all, in the occurrence and development of various diseases including inflammation and cancer, JMJD3 plays a crucial role, which has triggered a research boom among numerous scholars over the years. In this review, we primarily focused on the roles of JMJD3 in inflammation and cancers, and briefly discussed its application prospect, laying a theoretical foundation for further research and providing a train of thought for the prevention and treatment of related diseases.
Introduction
Histone modifications can alter structures and functions of genome, but the exact mechanisms remain enigmatic1. Histone methylation is dynamically regulated via methyltransferases and demethylases2. JmjC domain-containing protein D3 (JMJD3, also called KDM6B) is a member of the histone demethylase family, whose C-terminus with JmjC domain catalyzes demethylation, while the N-terminus with basic amino-acids clusters is responsible for nuclear placement3. JMJD3 and the ubiquitously transcribed X-chromosome tetratricopeptide repeat protein (UTX, also called KDM6A) have been identified to specifically demethylate H3K27me2/3, playing key roles in the epigenetic regulation of gene expression2. Seemingly, compared with UTX, JMJD3 is highly regulated at the transcriptional level and more susceptible to various stimuli like differentiation inducers and stress signals2.
The pivotal roles of JMJD3 and relevant mechanisms have been extensively studied for their involvement in cellular proliferation, differentiation, senescence and apoptosis4. From the perspective of tissue responses, JMJD3 mainly embodies in embryonic development, immune system, inflammation, neurodegenerative diseases and tumorigenesis5, 6. Herein, we primarily concentrate on the roles of JMJD3 in inflammation and cancer, aiming at laying a theoretical foundation for further research and providing a train of thought for the prevention and treatment of related diseases.
JMJD3 And Inflammation
The initiation, progression and termination of inflammation rely mostly on global activation of gene expression, post-transcriptional regulation, epigenetic modifications7. Undoubtedly, these processes are tightly controlled via specific signaling pathways, forming a criss-crossing network to finely regulate and balance all kinds of life activities, thereby avoiding inflammatory diseases. Although the inflammatory response varies from disease to disease, there is a series of transcription-activated genes that encode various mediators including inflammatory cytokines, adhesion molecules, chemokines, enzymes generating lipid mediators8.
As JMJD3 has emerged as a critical regulator of the macrophage and monocyte inflammation processes, the expression of JMJD3, generally low in tissues, can be strongly and quickly induced by various inflammatory stimuli and cellular stresses like lipopolysaccharide (LPS)6, 9. Recently, the mechanisms of JMJD3 in regulating the transcription of proinflammatory genes have been studied extensively and deeply, supporting the view that JMJD3 regulates gene expression in an enzymatic activity-dependent and -independent manner ?Figure 1?. According to research, in LPS-stimulated macrophages, the transcription of JMJD3 is dependent on multiple evolutionary conserved binding sites for the transcription factor NF-kB, after which other transcription factors such as ATF4, HIF-1, and the STAT family members like STAT1, STAT3, and STAT6 have been successively confirmed9-11. Consequently, JMJD3, as the downstream molecule of NF-kB, demethylates H3K27me3 to regulate the H3K27me3 levels and further controls transcriptional activity12. Moreover, studies suggest that newly synthesized JMJD3 is rapidly recruited to the transcription start sites (TSSs) of genes associated with basic H3K27me3 levels to enhance transcriptional initiation, those encoding LPS-inducible immune response and inflammatory mediators also included13. Puyu, et al. found that in human periodontal ligament cells (HPDLs) stimulated with LPS, the expression of JMJD3 was evidently increased and recruitment of JMJD3 to promoters of IL-6 and IL-12b enabled transcriptional activation in inflamed HPDLs by demethylation of H3K27me314. Similarly, in our previous studies, we observed that the nuclear transition and activation of NF-kB upregulated JMJD3 expression by LPS-stimulation in human umbilical vein endothelial cells (HUVECs). Besides, NF-kB/p65 and JMJD3 proteins were recruited to promoter regions of TNF-a, MMP-9, IL-1b, IL-6, TNF-a, and COX-2, where the H3K27me3 levels strikingly decreased, consistent with other related researches15, 16. Not only that, in promoter regions, JMJD3 helps to dissociate suppressive PRC2 (Polycomb Repressive Complex 2) responsible for methylation of H3K27, thus further facilitating the initiation of transcription6. Besides, after initiation, JMJD3, collaborated with histone demethylase KIAA1718 (also called KDM7A), promotes transcriptional elongation through expediting release of paused RNA polymerase II complex (RNAPII) and localizing elongation factors to target genes to effect RNAPII walking along the gene bodies rich in H3K27me317, 18. So, JMJD3 promotes expression of target genes by demethylating repressive H3K27me3 in both promoters and gene bodies, respectively modulating transcriptional initiation and elongation.
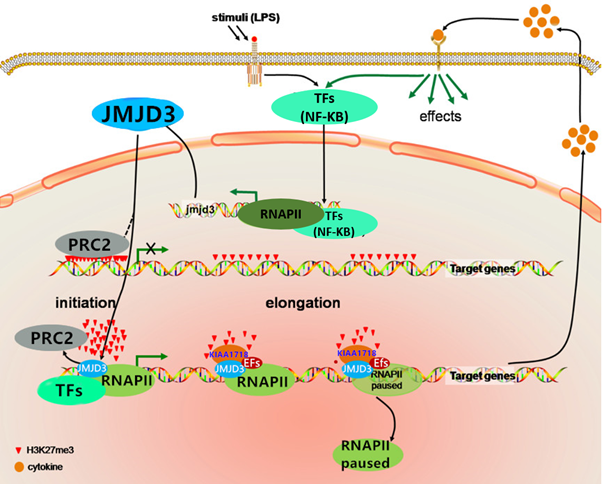
Figure 1: Model depicting the mechanisms of JMJD3 in the regulation of gene transcription. Activation of TFs leads to the expression of JMJD3, which drives the expression of target genes in an enzymatic activity-dependent and -independent manner.
In clinical diagnosis, elevated level of serum SAA is a signal of inflammatory diseases and is bound up with the pathogenesis of chronic diseases involving atherosclerosis, coronary heart disease, obesity, type 2 diabetes (T2D), rheumatoid arthritis (RA) and more19. And JMJD3 enhances the effects of SAA on foam cell formation by oxidized LDL, which plays a key role in the pathogenesis of these diseases especially atherosclerosise8, 19. For autoimmune-mediated inflammation like RA, JMJD3 in fibroblast-like synoviocytes (FLSs) was higher than healthy-FLSs, which by demethylation of H3K27me3, promoted the proliferation, invasion and migration of FLSs, contributing to joint inflammation, bone destruction and other pathologic processes20. Wu, et al. proposed that inhibiting the JMJD3 expression by the transcriptional factor Sp-1 or JMJD3-specific pharmacological inhibitor GSK-J4, helps to negatively regulate the inflammatory response and thus alleviates the progression of RA21. In atherosclerosis, research has revealed that JMJD3 causes a significant reduction in global H3K27me3 levels in vessels with advanced atherosclerotic plaques, underscoring the involvement of JMJD3 in the progression of atherosclerotic diseases22. Lately, however, Annette, et al. presented that myeloid JMJD3 deficiency enables not only the pathways involved in leukocyte chemotaxis to be obviously upregulated but atherosclerosis to progress to more advanced lesions with enhancement of collagen deposition and necrosis8. In T2D, JMJD3 by demethylase activity enhances H3K27me3-mediated repression at the promoter of IL-12, thus allowing for increased gene expression, which contributes to proinflammatory phenotype in bone marrow stem/progenitor cells23. Ulteriorly, researchers discovered in insulin-resistant and hyper-glycemic environment, JMJD3 along with up-regulated IL-12 arising therefrom can alter wound healing and may play a major role in sustained inflammation, also widely existing in other tissues like arterial walls and adipose tissues23. Based on these findings, we speculate that to some extent, JMJD3 exerts potential therapeutic effects on atherosclerosis, diabetic complications and other vascular inflammatory diseases, representing a novel option for ameliorating and treating related-disorders15.
JMJD3 And Cancer
As mentioned above, not only can JMJD3 regulate the expression of inflammatory factors via distinct signaling pathways, but can combine with promoters of cancer-related genes to exert dual effects on onset and progression of cancers. With different stages of inflammation-associated cancers, JMJD3 plays different roles in the disease progression. Typically, from the precancerous lesions to pancreatic cancers, the JMJD3 levels present a regular change-- from abnormal increase to gradual decrease with increased grade malignancy, revealing its significance in the pancreatic tumorigenesis24. In light of these findings, it’s not surprising that JMJD3 may act as a bridge between inflammation and cancer.
JMJD3 plays a crucial and dual role in cancer initiation and progression through binding to promoters of oncogenes or suppressor genes, demonstrating that either excess or lack of JMJD3 can have oncogenic effects25. In recent years, alteration of JMJD3 levels has emerged as a recurrent theme in many types of cancer. Moreover, in large number of clinical samples, aberrant expression of JMJD3 was commonly found, implying the far-reaching significance of research on JMJD3 to human final victory over cancers. Depending on cancer types, the expression levels and effects of JMJD3 are different (Table.1). Based on this table, we can roughly conclude that JMJD3 can promote most tumors and inhibit a few.
Table 1. Expression levels, effects and target genes of JMJD3 in various cancers
| Expression Level | Cancer Type | Effect | Target Genes (Signaling pathway) |
|---|---|---|---|
| High | Breast Cancer | Promoter | OCT4 (PHF20) , NANOG, SOX2, BCL226, 27 |
| Cervical Cancer | Promoter | HOX, p16INK4A(pRB-E2F)28 | |
| Ovarian Cancer | Promoter | HER2, MYCN29 | |
| Prostate Cancer | Promoter | TRA2A, ING3, FBXO11, CBX1, MYO1D, TENM4, GRIN3B MGMT, RPS6KA2, U2AF130, 31 | |
| Liver Cancer | Promoter | SLUG32 | |
| Pancreatic Ductal Adenocarcinoma | Suppressor | C/EBPα24 | |
| Renal Cell Carcinoma | Promoter | p16INK4A(pRB-E2F)33 | |
| Melanoma | Promoter | STC1, CCL2(NF-κB), BMP434 | |
| Medulloblastoma | Promoter | TopoIIβ35 | |
| Neuroblastoma | Promoter | BCL-236 | |
| T-ALL | Promoter | HES1, HEY1, NRARP(NOTCH1)37 | |
| AML | Promoter | HOX38 | |
| Hodgkin's lymphoma | Promoter | CD58, NOTCH2NL39 | |
| Diffuse Large B-Cell Lymphoma | Promoter | IRF4, BCL-2(NF-κB)40 | |
| Multiple Myeloma | Promoter | ELK1, FOS(MAPK, NF-κB)41 | |
| Low | Lung Cancer | Promoter | p2142 |
| Colorectal Cancer | Suppressor | p15INK4B43 |
With the deepening of research, a flurry of publications reported that carcinogenic and anti-cancer effects of JMJD3 involve its influences on proliferation, apoptosis, migration, stem cell behavior and tumor micro-environment. With respect to carcinogenic effects, the JMJD3 level, in most cases, is higher than that of normal people. Acute myeloid leukemia (AML) is a hematologic malignancy characterized by accumulation of undifferentiated myeloid precursors with resultant peripheral blood cytopenias44. In patients’ CD34+ stem cells with myelodysplastic syndrome (MDS, formerly known as preleukemia), the expression of JMJD3 abnormally increased. And high-risk patients are highly susceptible to AML5. Li, et al. found that the expression of JMJD3 was upregulated in AML cell lines. On the other hand, by applying GSK-J4, they observed that the expression of HOX genes, DNA replication and cell-cycle relevant pathways were significantly downregulated. Meanwhile, H3K27me3 was obviously increased in the TSSs of HOX genes, conversely suggesting JMJD3 can upregulate the expression of the oncogene HOX by demethylation H3K27 to promote cancer38. Furthermore, the levels of HOX gene expression were positively correlated with the mRNA levels of JMJD3 among AML patients45. In T-cell acute lymphoblastic leukemia (T-ALL), JMJD3, as part of the NOTCH1 transcriptional complex, plays an indispensable role in initiation and maintenance of the blood cancer. Interestingly, JMJD3 and UTX, both as demethylase, yet have the opposite effects. JMJD3 control oncogenic gene targets like NOTCH1 and MYC, while UTX acts as a suppressor37.
In contrast, as for anticarcinogenic effect, usually, the expression of JMJD3 is inhibited. Few as tumor types are in number, the significant meaning of JMJD3 can’t be neglected as well, whether in scientific research or clinical practice. In colorectal cancer (CRC) tissue, JMJD3 expression levels along with vitamin D receptors (VDRs) are significantly lower than normal tissues. Reportedly, in CRC patients, JMJD3 was induced by the vitamin D metabolite 1α,25-dihydroxyvitamin D3 (1,25(OH)2D3), which suppressed epithelial-to-mesenchymal transition (EMT), a major event in epithelial cancers46. In addition, through knock-down (KD) experiments of JMJD3, researchers found that KD-JMJD3 suppressed the expression of cell-cycle regulator P15INK4B, which encodes transcriptional inhibitors of cyclin-dependent kinases43. As a result, KD-JMJD3 promoted cell-cycle progression and suppressed apoptosis, forcefully corroborating anticarcinogenic roles of JMJD343.
Notably, though, JMJD3 appears to play a dual role in some tumors, attracting increasing attention. For example, most patients with Renal Cell Carcinoma (RCC), whose JMJD3 expression is upregulated, yet have a better prognosis33. Presumably, the discrepancy may lie in different microenvironments, being at an early stage of cancer, and other unknown functions of JMJD3, requiring further elucidation.
Future Prospects
The identification of JMJD3 has opened a novel frontier in the study of dynamic epigenetic regulation, although mechanically, the post-translational modifications themselves are chemically stable22. Beyond doubt, JMJD3 is widely implicated in distinct signaling pathways such as NF-κB, STATs, and there exist intricate interrelationships, sparking the upsurge of research on its upstream regulatory signals, networks as well as downstream molecular effects and mechanisms. The involvement of JMJD3 in diseases including inflammation and cancer also provides a unique opportunity for pharmacological intervention by seeking small molecule inhibitors to counteract their function. Indeed, the inhibitors like GSK-J4 have been widely used in scientific studies. Researches on GSK-J4 treatment of K27M-expression cells from patients with pediatric brainstem glioma, have presented us a dose-dependent inhibition of survival, offering a valid therapeutic strategy for pediatric brainstem glioma47. Recently, Yunan, et al. revealed that GSK-J4 has a therapeutic potential for patients with AML38. Our another study suggested that in LPS-stimulated macrophages, NF-kB signaling pathway was blocked by lanthanum, a light rare earth element via inhibiting inflammatory gene expression. Subsequently, in LPS-activated HUVECs, we found that lanthanum chloride inhibited the activation of NF-kB as well as downstream cytokines and adhesion molecules, where the effect of JMJD3 was quite pivotal48, 49. Given the broad spectrum of genes regulated by JMJD3 plus the complexity of the mechanism, further in-depth researches remain to be quite necessary. Absolutely, understanding how JMJD3 participates in various cellular processes, is of great significance in advancing our understanding of multiple roles and mechanisms of JMJD3 in related clinical diseases, laying a theoretical basis for future research and holding great promise for drug screening and therapeutic intervention.
Acknowledgement
This article was published by financial support of grants from the National Natural Science Foundation of China (grant number 81772083, 81860342).
References
-
- Völker-Albert MC, Schmidt A, Forne I, et al. Analysis of Histone Modifications by Mass Spectrometry. Curr Protoc Protein Sci. 2018; 92(1): e54.
- Hong S, Cho YW, Yu LR, et al. Identification of JmjC domain-containing UTX and JMJD3 as histone H3 lysine 27 demethylases. Proc Natl Acad Sci U S A. 2007; 104(47): 18439-44.
- Kamikawa YF, Donohoe ME. The localization of histone H3K27me3 demethylase Jmjd3 is dynamically regulated. Epigenetics. 2014; 9(6): 834-41.
- Yang D, Yu B, Sun H, et al. The Roles of Histone Demethylase Jmjd3 in Osteoblast Differentiation and Apoptosis. J Clin Med. 2017; 6(3).
- Burchfield JS, Li Q, Wang HY, et al. JMJD3 as an epigenetic regulator in development and disease. Int J Biochem Cell Biol. 2015; 67: 148-57.
- Salminen A, Kaarniranta K, Hiltunen M, et al. Histone demethylase Jumonji D3 (JMJD3/KDM6B) at the nexus of epigenetic regulation of inflammation and the aging process. J Mol Med (Berl). 2014; 92(10): 1035-43.
- Nicodeme E, Jeffrey KL, Schaefer U, et al. Suppression of inflammation by a synthetic histone mimic. Nature. 2010; 468(7327): 1119-23.
- Neele AE, Gijbels MJJ, van der Velden S, et al. Myeloid Kdm6b deficiency results in advanced atherosclerosis. Atherosclerosis. 2018; 275: 156-165.
- Lee HY, Choi K, Oh H, et al. HIF-1-dependent induction of Jumonji domain-containing protein (JMJD) 3 under hypoxic conditions. Mol Cells. 2014; 37(1): 43-50.
- Penas C, Navarro X. Epigenetic Modifications Associated to Neuroinflammation and Neuropathic Pain After Neural Trauma. Front Cell Neurosci. 2018; 12: 158.
- Przanowski P, Dabrowski M, Ellert-Miklaszewska A, et al. The signal transducers Stat1 and Stat3 and their novel target Jmjd3 drive the expression of inflammatory genes in microglia. J Mol Med (Berl). 2014; 92(3): 239-54.
- De Santa F, Totaro MG, Prosperini E, et al. The Histone H3 Lysine-27 Demethylase Jmjd3 Links Inflammation to Inhibition of Polycomb-Mediated Gene Silencing. Cell. 2007; 130(6): 1083-1094.
- De Santa F, Narang V, Yap ZH, et al. Jmjd3 contributes to the control of gene expression in LPS-activated macrophages. EMBO J. 2009; 28(21): 3341-52.
- Wang P, Yue J, Xu W, et al. Jumonji domain-containing protein 3 regulates the early inflammatory response epigenetically in human periodontal ligament cells. Archives of Oral Biology. 2018; 93: 87-94.
- Chen X, Xiu M, Xing J, et al. Lanthanum Chloride Inhibits LPS Mediated Expressions of Pro-Inflammatory Cytokines and Adhesion Molecules in HUVECs: Involvement of NF-κB-Jmjd3 Signaling. Cellular Physiology and Biochemistry. 2017; 42(5): 1713-1724.
- Yu S, Chen X, Xiu M, et al. The regulation of Jmjd3 upon the expression of NF-kappaB downstream inflammatory genes in LPS activated vascular endothelial cells. Biochem Biophys Res Commun. 2017; 485(1): 62-68.
- Chen S, Ma J, Wu F, et al. The histone H3 Lys 27 demethylase JMJD3 regulates gene expression by impacting transcriptional elongation. Genes Dev. 2012; 26(12): 1364-75.
- Estarás C, Fueyo R, Akizu N, et al. RNA polymerase II progression through H3K27me3-enriched gene bodies requires JMJD3 histone demethylase. Mol Biol Cell. 2013; 24(3): 351-60.
- Yan Q, Sun L, Zhu Z, et al. Jmjd3-mediated epigenetic regulation of inflammatory cytokine gene expression in serum amyloid A-stimulated macrophages. Cell Signal. 2014; 26(9): 1783-91.
- Jia W, Wu W, Yang D, et al. Histone demethylase JMJD3 regulates fibroblast-like synoviocyte-mediated proliferation and joint destruction in rheumatoid arthritis. FASEB J. 2018; 32(7): 4031-4042.
- Wu W, Qin M, Jia W, et al. Cystathionine-gamma-lyase ameliorates the histone demethylase JMJD3-mediated autoimmune response in rheumatoid arthritis. Cell Mol Immunol. 2018.
- Wierda RJ, Rietveld IM, van Eggermond MC, et al. Global histone H3 lysine 27 triple methylation levels are reduced in vessels with advanced atherosclerotic plaques. Life Sci. 2015; 129: 3-9.
- Gallagher KA, Joshi A, Carson WF, et al. Epigenetic changes in bone marrow progenitor cells influence the inflammatory phenotype and alter wound healing in type 2 diabetes. Diabetes. 2015; 64(4): 1420-30.
- Yamamoto K, Tateishi K, Kudo Y, et al. Loss of histone demethylase KDM6B enhances aggressiveness of pancreatic cancer through downregulation of C/EBPalpha. Carcinogenesis. 2014; 35(11): 2404-14.
- Agger K, Cloos PA, Christensen J, et al. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature. 2007; 449(7163): 731-4.
- Yan N, Xu L, Wu X, et al. GSKJ4, an H3K27me3 demethylase inhibitor, effectively suppresses the breast cancer stem cells. Experimental Cell Research. 2017; 359(2): 405-414.
- Xun J, Wang D, Shen L, et al. JMJD3 suppresses stem cell-like characteristics in breast cancer cells by downregulation of Oct4 independently of its demethylase activity. Oncotarget. 2017; 8(13): 21918-21929.
- McLaughlin-Drubin ME, Park D, Munger K. Tumor suppressor p16INK4A is necessary for survival of cervical carcinoma cell lines. Proc Natl Acad Sci U S A. 2013; 110(40): 16175-80.
- Mo J, Wang L, Huang X, et al. Multifunctional nanoparticles for co-delivery of paclitaxel and carboplatin against ovarian cancer by inactivating the JMJD3-HER2 axis. Nanoscale. 2017; 9(35): 13142-13152.
- Daures M, Idrissou M, Judes G, et al. A new metabolic gene signature in prostate cancer regulated by JMJD3 and EZH2. Oncotarget. 2018; 9(34): 23413-23425.
- Ngollo M, Lebert A, Daures M, et al. Global analysis of H3K27me3 as an epigenetic marker in prostate cancer progression. BMC Cancer. 2017; 17(1): 261.
- Tang B, Qi G, Tang F, et al. Aberrant JMJD3 Expression Upregulates Slug to Promote Migration, Invasion, and Stem Cell-Like Behaviors in Hepatocellular Carcinoma. Cancer Res. 2016; 76(22): 6520-6532.
- Shen Y, Guo X, Wang Y, et al. Expression and significance of histone H3K27 demethylases in renal cell carcinoma. BMC Cancer. 2012; 12: 470.
- Park WY, Hong BJ, Lee J, et al. H3K27 Demethylase JMJD3 Employs the NF-kappaB and BMP Signaling Pathways to Modulate the Tumor Microenvironment and Promote Melanoma Progression and Metastasis. Cancer Res. 2016; 76(1): 161-70.
- Chen J, Zhao J, Zhou X, et al. Immunohistochemical investigation of topoIIbeta, H3K27me3 and JMJD3 expressions in medulloblastoma. Pathol Res Pract. 2017; 213(8): 975-981.
- Lochmann TL, Powell KM, Ham J, et al. Targeted inhibition of histone H3K27 demethylation is effective in high-risk neuroblastoma. Sci Transl Med. 2018; 10(441).
- Ntziachristos P, Tsirigos A, Welstead GG, et al. Contrasting roles of histone 3 lysine 27 demethylases in acute lymphoblastic leukaemia. Nature. 2014; 514(7523): 513-7.
- Li Y, Zhang M, Sheng M, et al. Therapeutic potential of GSK-J4, a histone demethylase KDM6B/JMJD3 inhibitor, for acute myeloid leukemia. J Cancer Res Clin Oncol. 2018; 144(6): 1065-1077.
- Anderton JA, Bose S, Vockerodt M, et al. The H3K27me3 demethylase, KDM6B, is induced by Epstein-Barr virus and over-expressed in Hodgkin's Lymphoma. Oncogene. 2011; 30(17): 2037-43.
- Zhang Y, Shen L, Stupack DG, et al. JMJD3 promotes survival of diffuse large B-cell lymphoma subtypes via distinct mechanisms. Oncotarget. 2016; 7(20): 29387-99.
- Ohguchi H, Harada T, Sagawa M, et al. KDM6B modulates MAPK pathway mediating multiple myeloma cell growth and survival. Leukemia. 2017; 31(12): 2661-2669.
- Tian C, Deng H, Tang X, et al. [Effect of Jumonji domain-containing protein-3 on the proliferation and migration of lung cancer cell line]. Sheng Wu Yi Xue Gong Cheng Xue Za Zhi. 2012; 29(3): 514-8.
- Tokunaga R, Sakamoto Y, Nakagawa S, et al. The Prognostic Significance of Histone Lysine Demethylase JMJD3/KDM6B in Colorectal Cancer. Ann Surg Oncol. 2016; 23(2): 678-85.
- Papaemmanuil E, Gerstung M, Bullinger L, et al. Genomic Classification and Prognosis in Acute Myeloid Leukemia. N Engl J Med. 2016; 374(23): 2209-21.
- Skvarova Kramarzova K, Fiser K, Mejstrikova E, et al. Homeobox gene expression in acute myeloid leukemia is linked to typical underlying molecular aberrations. J Hematol Oncol. 2014; 7: 94.
- Pereira F, Barbáchano A, Silva J, et al. KDM6B/JMJD3 histone demethylase is induced by vitamin D and modulates its effects in colon cancer cells. Hum Mol Genet. 2011; 20(23): 4655-65.
- Hashizume R, Andor N, Ihara Y, et al. Pharmacologic inhibition of histone demethylation as a therapy for pediatric brainstem glioma. Nat Med. 2014; 20(12): 1394-6.
- Guo F, Lou Y, Feng N, et al. Exposure to lanthanum compound diminishes LPS-induced inflammation-associated gene expression: involvements of PKC and NF-κB signaling pathways. BioMetals. 2010; 23(4): 669-680.
- Guo F, Guo X, Xie A, et al. The Suppressive Effects of Lanthanum on the Production of Inflammatory Mediators in Mice Challenged by LPS. Biological Trace Element Research. 2011; 142(3): 693-703.
